Theme n°1 : ABC-F translation factors
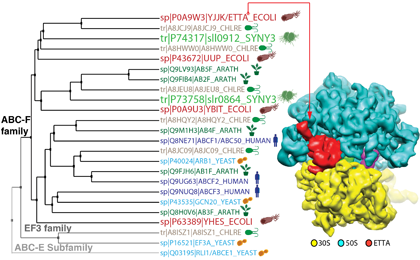
The ABC-F family is a family of soluble protein, which belongs to the larger ABC (ATP Binding Cassette) protein superfamily. This superfamily comprises ATPase proteins formed by 2 homologous ABC domains. Most of the ABC proteins are membrane transporters, but the ABC-F family is characterized by soluble proteins. It is the most widespread family of soluble proteins within the ABC superfamily. Four representatives of this ABC-F family are present in Escherichia coli, two in the cyanobacterium Synechocystis, six in Chlamydomonas reinhardtii (including one chloroplastic), five in plants, and three in humans. With our collaborators, we have demonstrated that the most abundant ABC-F family member found in eubacteria, the protein EttA (energy-sensing translational throttle A), is a protein translation factor that gates ribosome entry into the elongation cycle in an ADP-to-ATP ratio-dependent manner.
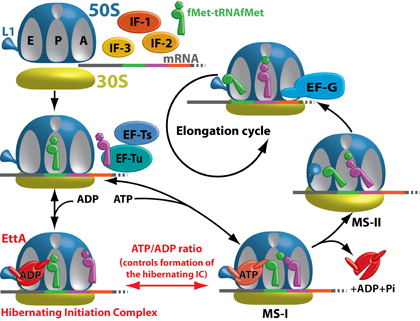
In the presence of ADP, EttA inhibits formation of the first peptide bond by the 70S Initiation Complex, thereby blocking entry into the translation elongation cycle. The ATP-bound conformation of EttA stabilizes the 70S Initiation Complex in a conformation favorable to the formation of the first peptide bond. This interaction stimulates ATP hydrolysis by EttA, leading its dissociation from the ribosome, thereby allowing EF-G to catalyze the first round of translocation and drive entry of the 70S ribosomal complex into the elongation cycle. In vivo, cells with the ettA gene knocked out, showed a severe fitness defect in long-term stationary phase, a condition in which the intracellular ATP level is low.
Since translational apparatus consumes more than half of the total cellular energy and it is thus not unexpected to find a new translation factor that regulates this process. We are currently extending our study of EttA to three other paralogs present in E. coli and ortholog of EttA responsible for antibiotic resistance in several pathogenic bacteria. We also are investigating the function of the two ABC-F proteins present in the cyanobacterium Synechocystis. Our goal is to map the different actions of the ABC-F proteins on the translation apparatus.
Theme n°2 : Functionnal characterization of novel translation factors
In this second task we will develop a screening methodology to identify the phenotype related to translation defects in a mutant collection. This collection will be composed of deletion mutants of proteins known to interact with the ribosome and for which the function remains unclear. In parallel, we will attempt a pull-down approach to identify new factors that interact with stalled ribosomes.
